|
When multiple strands from different regions of a polypeptide interact in this way Figure 2 The crystal structure of a synthetic glycine polymer suggested that folded proteins feature hydrogen bonds between backbone. ABSTRACT The polypeptide backbone fold in the solution structure of murine epidermal growth factor has been deter- mined by nuclear magnetic resonance. As few as two strands can interact in this way. The C-O and C-N distances are related but their trends are not strictly connected to peptide-bond planarity, although a correlation amongst all of these parameters is expected on the basis of the classical resonance model. Instead, hydrogen bonds form between the backbone atoms of separate -strands. The R groups are attached to the carbons and extend above and below the pleats folds. In the -pleated sheet, hydrogen bonding between atoms on the polypeptide chains backbone form the 'pleats'. The polypeptides R groups (the variant groups) protrude out from the -helix chain. On the other hand, the variability of the C-O and C-N distances is related to the strength of the interactions between the lone pair of the N atom and the C-O π* system, which is modulated by the ψ angle. The structure of cell-surface mucins consists of an unstructured central polypeptide backbone with densely grafted polymeric sugar side chains (glycans) and a transmembrane domain that directly. Every helical turn in an alpha helix has 3.6 amino acid residues. 
Maximum and minimum values for each bond distance are found for conformations with the specific bond perpendicular and parallel to the adjacent CONH peptide plane, respectively. Proteins are polypeptide structures consisting of one or more long chains of amino acid residues. The 'polypeptide backbone' refers to all atoms in a polypeptide chain, except for those that form the peptide bonds. Ribosomes catalyses the peptide bond formation. Amino group is the beginning of the polypeptide, and the carboxyl group is at the end. Generally, the total number of nonpolar amino acids has a greater effect on protein structure than the exact order of amino acids in a polypeptide chain. 2 Amino acids join, a water molecule is lost in a condensation reaction, and a peptide group is created. The dependence of C α distances on ψ is governed by interactions between the σ system of the C α moiety and the C-O π system of the peptide bond. If a statement is false, explain why it is false. The overall agreement between the computed and the statistical data suggests that these trends are essentially driven by local effects. 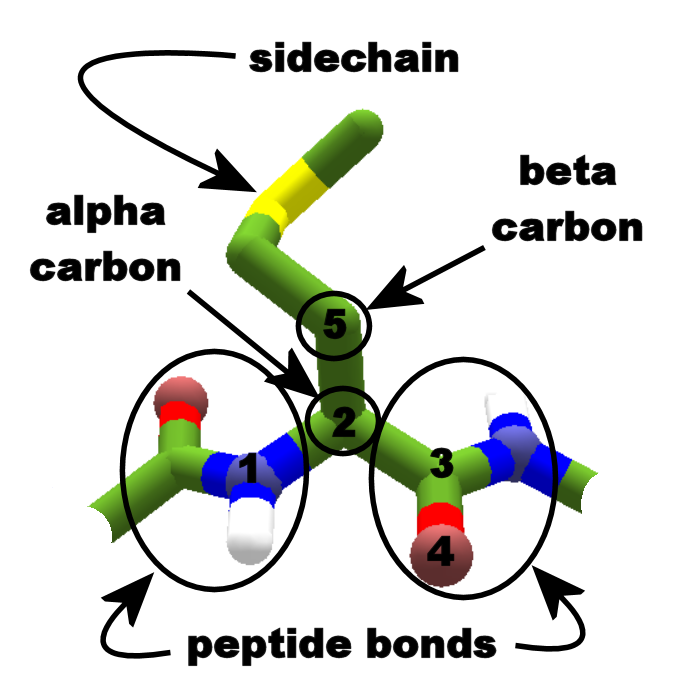
All of these bond lengths indeed display a systematic variability in the ψ angle according to both calculations and surveys of protein structures. Download scientific diagram Electronic configuration of the polypeptide backbone and conformational behavior of A proteins. polypeptides backbone (main chain) atoms tertiary structure refers to the three-dimensional structure of an entire polypeptide chain and quaternary. \): Tertiary structure of Carboxypeptidase A, from bovine pancreas, showing \(\alpha\)-helix (red), \(\beta\)-pleated sheets (yellow) and or random coil (green).By combining quantum-mechanical analysis of small model peptides and statistical surveys of high-resolution protein structures, a systematic conformational dependence of bond lengths in polypeptide backbones has been unveiled which involves both the peptide bond (C-O and C-N) and those bonds centred on the C α atom.
0 Comments
Leave a Reply. |
AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |
 RSS Feed
RSS Feed
